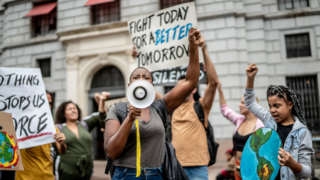USC Today Home Page
Top Stories from USC News
Dozens of student-focused events will recognize the newest graduates.
The clinic is the first in Nevada to offer in-state heart transplant services.
USC alum Chien, who recently starred in The Brothers Sun on Netflix, is a master of combining storytelling and choreography.
Bush had a legendary career at USC and an electrifying 2005 season, scoring 18 touchdowns.

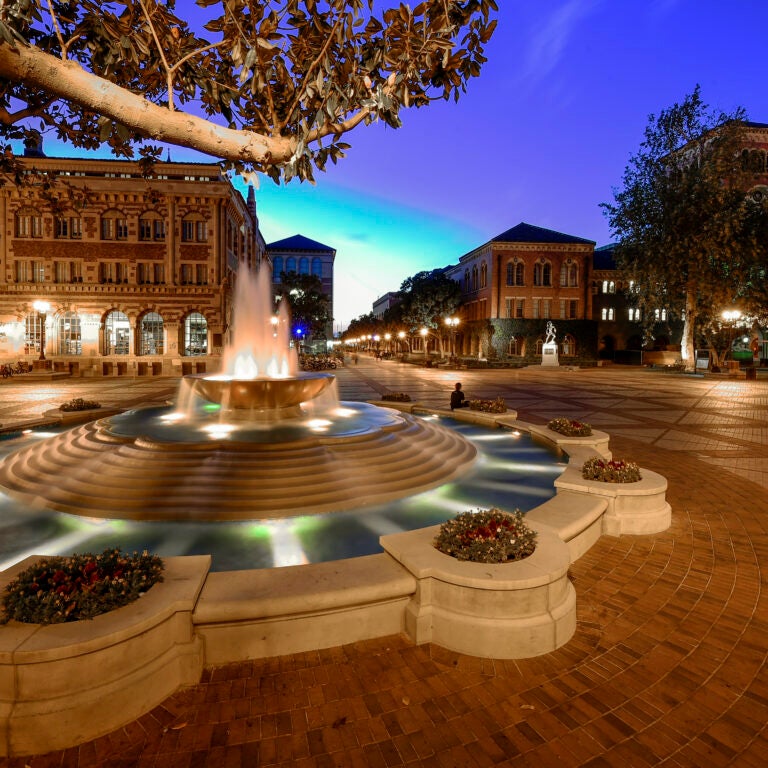
Assignment: Earth begins with us
Assignment: Earth is USC’s comprehensive sustainability framework, which guides the university in its mission to foster a healthy, just, and thriving campus and global community. It encompasses various initiatives and strategies to promote environmental stewardship, social equity and economic vitality within the university and beyond.
Celebrating Earth Month at USC
Celebrating Earth Month at USC includes events, initiatives and activities dedicated to raising awareness about environmental issues, promoting sustainability practices and inspiring action among the university community. Here are our latest stories.
Eco Film & Media Arts Festival, Urban Trees Initiative receive national sustainability accolades
USC programs are recognized for student engagement and racial equity from the Association for the Advancement of Sustainability in Higher Education.
Three USC schools collaborate to reimagine the L.A. River
Researchers in landscape architecture, fluid mechanics and augmented reality have created an interactive hydraulic model alongside the river.
USC Sea Grant research, community outreach show approval of aquaculture can be earned
People across all demographics could change their minds when they learned about the environmental benefits of seafood farming, USC research suggests.
From throwing soup to suing governments, there’s strategy to climate activism’s seeming chaos
USC’s Shannon Gibson says that with international climate talks failing to make progress fast enough, activists are radically rethinking how to be most effective in the streets, political arenas and courtrooms.
Surprising discovery about coral’s resilience could help reefs survive climate change
USC Dornsife researchers studying a common Caribbean coral’s ability to adapt to rising temperatures turn up an unexpected result.

Looking to the Sky
Trojans gathered in Hahn Plaza on the University Park Campus on Monday for a lively eclipse viewing party. Learn from USC experts about the symbolism of the celestial rarity and more.